Contact
 Professor Hans Ronne
Professor Hans Ronne
Department of Forest Mycology and Plant Pathology
018-673223, hans.ronne@slu.se

The moss Physcomitrella is a small plant that has become an important tool in plant research. The reason for this is that it is easy to inactivate (knock out) genes in this moss by using homologous recombination, so called reverse genetics.
This is something that has long been possible in model organisms such as yeast and mice, but not in plants. The point of knocking out genes is that it makes it possible to figure out their functions by studying the properties of individuals that lack the gene.
Physcomitrella has therefore developed into an important model plant. The sequence of the Physcomitrella genome has been determined, which has further increased its value as a model organism.
Another important tool in molecular genetics is plasmids, small DNA molecules that can replicate inside the cells of the host organism. Plasmids make it possible to clone new genes, and to study the function of already cloned genes by overexpression. The latter is possible because plasmids frequently exists in many copies per cell and thus produce more proteins than the genes of the host cell, which exist in only one or two copies.
The use of plasmids has so far for practical reasons been limited to unicellular organisms such as bacteria and yeast. Plasmids can replicate also in plant and animal cells, but these plasmids are frequently rearranged, and insert themselves into the host cell genome, which makes them much more difficult to use.
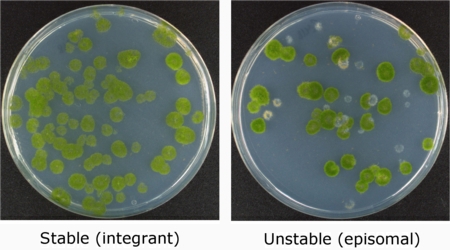
However, we have found that plasmids that replicate in Physcomitrella are stable and can be rescued back into bacteria without rearrangements (Murén et al., 2009). This makes it possible to build so called shuttle plasmids that can be used to move genes back and forth between moss, bacteria and yeast.
We are now following up this discovery by developing and testing new powerful methods that use plasmids in moss. These methods come from yeast, where plasmids have been used with great success for many years.
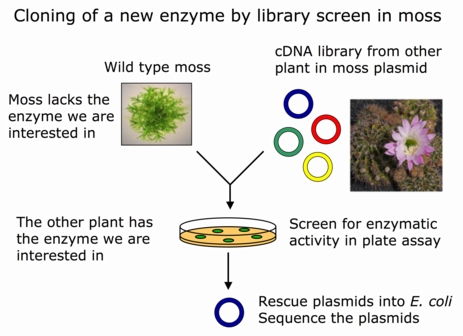
One such application is cloning of new genes from plasmid libraries where DNA from other plants has been inserted into a moss plasmid. It is already today possible to do such cloning in yeast, but since yeast is not a plant, genes needed for photosynthesis and other plant specific processes do not work in yeast. By using moss instead, it will be easier to clone and study plant genes.
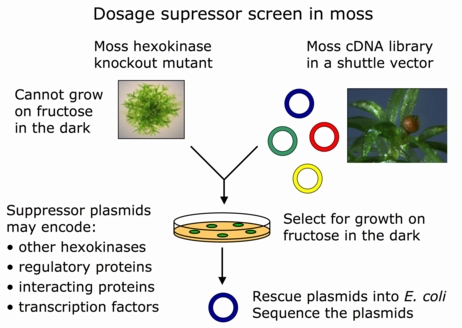
Another application is gene dosage suppression, where a plasmid library is screened for genes that can suppress the effect of a given mutation when overexpressed from a plasmid. This makes it possible to quickly find new genes that are functionally related to the gene of interest. It is frequently possible to unravel an entire network of genes that function together in a given biological process by using dosage suppression.
These methods will be useful both in basic research and in biotechnology and crop improvement. Dosage suppression will make it possible to quickly discover the function of new plant genes. By using moss plasmids it will also be possible to clone and study plant genes that do not work in yeast. This will be particularly useful in plant biotechnology, where you frequently want to clone genes for enzymes that are unique to plants.
Nylander, M., Heino, P., Helenius, E., Palva, T., Ronne, H. and Welin, B. (2001). The low temperature- and salt-induced RCI2Agene of Arabidopsis complements the sodium sensitivity caused by a deletion of the homologous yeast gene SNA1. Plant Mol. Biol. 45, 341-352.
Lindroth, A., Saarikoski, P., Flygh, G., Clapham, D., Grönroos, R. Thelander, M., Ronne, H. and von Arnold, S. (2001). Two S-adenosylmethionone synthase genes that are differentially expressed during adventitious root development in Pinus contorta. Plant Mol. Biol. 46, 336-346.
Thelander, M., Fredriksson, D., Schouten, J., Hoge, H. and Ronne, H. (2002). Cloning by pathway activation in yeast: identification of an Arabidopsis thaliana F-box protein that turns on glucose repression. Plant Mol. Biol. 49, 67-79.
Olsson, T., Thelander, M., and Ronne, H. (2003) A novel type of chloroplast stromal hexokinase is the major glucose phosphorylating enzyme in the moss Physcomitrella patens. J. Biol. Chem. 278, 44439-44447.
Thelander, M., Olsson, T., and Ronne, H. (2004) Snf1-related protein kinase 1 is needed for growth in a normal day-night light cycle. EMBO J. 23, 1900–1910.
Thelander, M., Olsson, T., and Ronne, H. (2005) Evidence that the balance between chloronemal and caulonemal growth is regulated by the energy supply in the moss Physcomitrella patens. J. Exp. Bot. 56, 653-662.
Thelander, M., Nilsson, A., Olsson, T., Johansson, M., Girod, P.-A., Schaefer, D.G., Zrÿd, J.P. and Ronne, H. (2007) The moss genes PpSKI1 and PpSKI2 encode nuclear SnRK1 interacting proteins with homologues in vascular plants. Plant Mol. Biol. 64, 559-573.
Thelander, M., Nilsson, A. and Ronne, H. (2009) Carbon and energy metabolism. In The Moss Physcomitrella, Eds. Perroud P.-F., Knight, C. and Cove, D., Annual Plant Reviews 36, 211-245.
Murén, E., Nilsson, A., Ulfstedt, M., Johansson, M. and Ronne, H. (2009) Rescue and characterization of episomally replicating DNA from the moss Physcomitrella. Proc. Natl. Acad. Sci. USA 106, 19444-19449.
Eklund, D.M., Thelander, M., Landberg, K., Ståldahl, V., Nilsson, A., Johansson, M., Valsecchi, I., Pederson, E., Kowalczyk, M., Ljung, K., Ronne, H. and Sundberg, E. (2010) Physcomitrella patens homologues of the Arabidopsis thaliana SHI/STY/LRP genes control auxin biosynthesis and affect growth and development in moss. Development 137, 1275-1284.
Nilsson, A., Olsson, T., Ulfstedt, M., Thelander, M. and Ronne, H. (2011) Two novel types of hexokinases in the moss Physcomitrella patens. BMC Plant Biology 11: 32.
Elfving, N., Davoine, C., Benlloch, R., Blomberg, J., Brännström, K., Müller, D., Nilsson, A., Ulfstedt, M., Ronne, H., Wingsle, G., Nilsson, O. and Björklund, S. (2011) The Arabidopsis thaliana Med25 mediator subunit integrates environmental cues to control plant development. Proc. Natl. Acad. Sci. USA 108, 8245-8250.
Hörnblad, E., Ulfstedt, M., Ronne, H. and Marchant, A. (2013) Partial functional conservation of IRX10 homologs in Physcomitrella patens and Arabidopsis thaliana indicates an evolutionary step contributing to vascular formation in land plants. BMC Plant Biology 13: 3.
Lind C, Dreyer I, López-Sanjurjo EJ, von Meyer K, Ishizaki K, Kohchi T, Lang D, Zhao Y, Kreuzer I, Al-Rasheid KA, Ronne H, Reski R, Zhu JK, Geiger D, Hedrich R (2015) Stomatal guard cells co-opted an ancient ABA-dependent desiccation survival system to regulate stomatal closure. Curr. Biol. 25, 928-935.
Ulfstedt, M., Hu, G.-Z., Johansson, M., and Ronne, H. (2017) Testing of Auxotrophic Selection Markers for Use in the Moss Physcomitrella Provides New Insights into the Mechanisms of Targeted Recombination. Front. Plant Sci. 8: 1850.
 Professor Hans Ronne
Professor Hans RonneDepartment of Forest Mycology and Plant Pathology
018-673223, hans.ronne@slu.se