Facts:
- Funding: FORMAS
- Funding period: 2019-2021
The aim of this project is to increase the understanding of the formation of methylmercury, a toxic form of mercury, and the microbes mediating this process in the Baltic Sea.
Methylmercury (MeHg), a neurotoxic compound biomagnifying through aquatic food webs, can be a threat to human health via fish consumption. However, the composition and distribution of the microbial communities mediating the methylation of mercury (Hg) to MeHg in marine systems remain largely unknown. The aim of this project is to fill this knowledge gap.
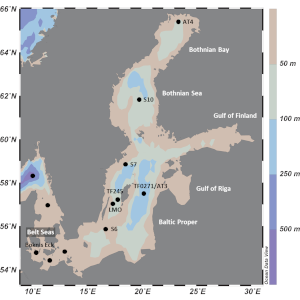
Analyses of metagenomes (genetic material recovered from environmental samples) and metatranscriptomes (gene expression of entire microbial communities) obtained from water and sediment samples collected in the Baltic Sea will be done to determined the distribution and activity of the genes (hgcAB genes) involved in Hg methylation, as well as the identity of the microbial taxa carrying the genes.
In both pelagic redoxclines (a layer of water with a strong vertical gradient between the upper oxygenated and lower anoxic water) and sediment, the distribution and activity of hgcAB-carrying microbes is found to differ related to oxygen concentrations. In both environments, the phylogenetic analysis identifies well-characterized Hg methylators, mainly within Deltaproteobacteria, but also undescribed Hg methylators e.g., members of Kiritimatiellaeota/Verrucomicrobia. Transcripts from unidentified HgcAB-carrying microbes are also detected in oxygen-deficient water and sediments illustrating the lack of knowledge related to Hg methylators in environments.
Altogether, the project aims at unveiling the diversity and activity of the microorganisms responsible for the formation of MeHg in the Baltic Sea and pinpoint the ecological niche of these microorganisms within the marine water column and oxygen-deficient sediments.